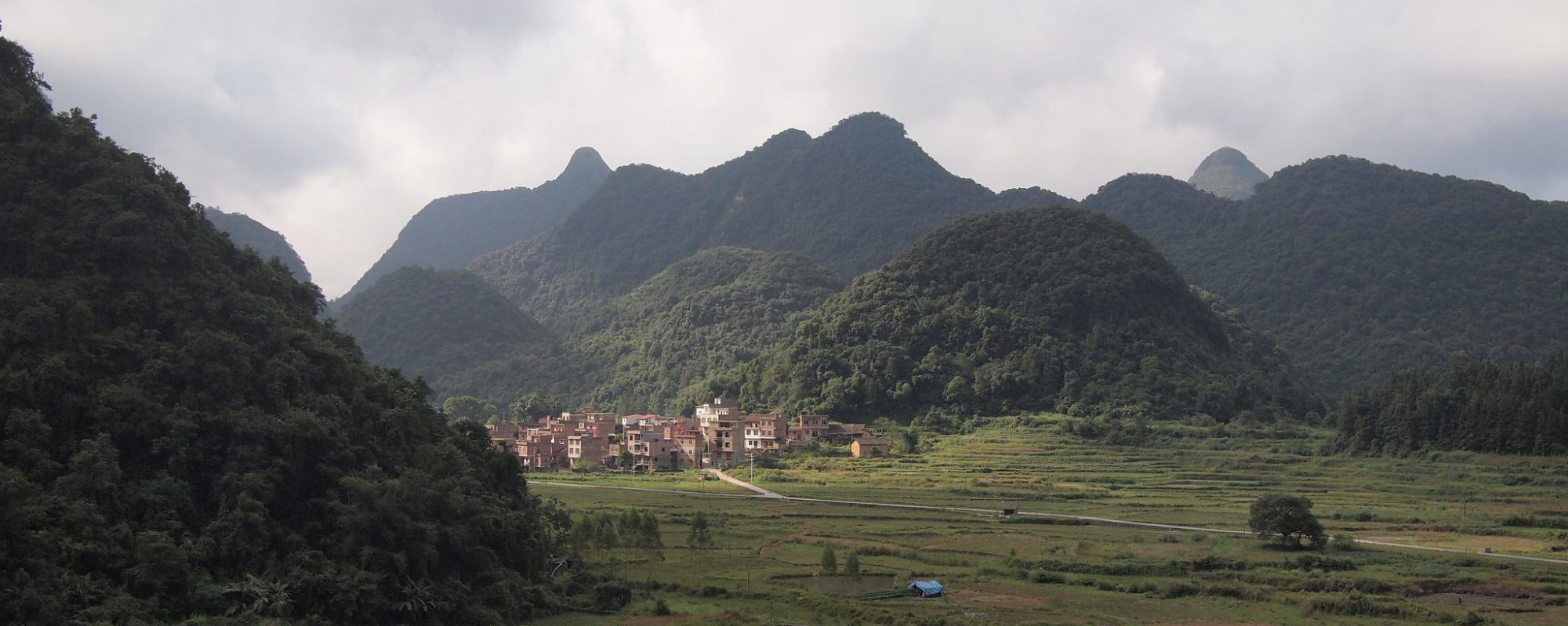
Coarse scaled NDVI (or FAPAR, EVI...) images have a high temporal frequency and are delivered as Maximum Value Composites (MVC) of several days, which means the highest value is taken, assuming that clouds and other contaminated pixels have low values. However, especially in areas with a rainy season, the composites over 10-16 days still contain … Continue reading Smoothing/Filtering a NDVI time series using a Savitzky Golay filter and R
Tag: R
Pixel-wise regression between two raster time series (e.g. NDVI and rainfall)
Doing a pixel-wise regression between two raster time series can be useful for several reasons, for example: find the relation between vegetation and rainfall for each pixel, e.g. a low correlation could be a sign of degradation derive regression coefficients to model the depending variable using the independend variable (e.g. model NDVI with rainfall data) … Continue reading Pixel-wise regression between two raster time series (e.g. NDVI and rainfall)
Pixel-wise time series trend anaylsis with NDVI (GIMMS) and R
The GIMMS dataset is currently offline and the new GIMMS3g will soon be released, but it does not really matter which type of data is used for this purpose. It can also be SPOT VGT, MODIS or anything else as long as the temporal resolution is high and the time frame is long enough to … Continue reading Pixel-wise time series trend anaylsis with NDVI (GIMMS) and R
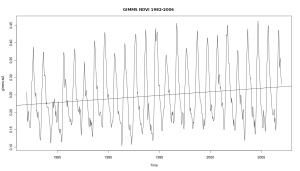
Simple time series analysis with GIMMS NDVI and R
Time series analysis with satellite derived greenness indexes (e.g. NDVI) is a powerfull tool to assess environmental processes. AVHRR, MODIS and SPOT VGT provide global and daily imagery. Creating some plots is a simple task, and here is a rough start how it is done with GIMMS NDVI. All we need is the free software … Continue reading Simple time series analysis with GIMMS NDVI and R
Converting GPCC gridded rainfall 1901-2010@0.5° to monthly Geotiffs
GPCC rainfall data contains the largest database worldwide, with over 90 000 weather-stations interpolated to a 0.5° grid. It includes S. Nicholsons dataset for Africa and is thus a good source for gridded monthly precipitation. It covers the period 1901-2010, however, it comes in a strange dataformat and it is a long way until we get … Continue reading Converting GPCC gridded rainfall 1901-2010@0.5° to monthly Geotiffs
Automatically downloading and processing TRMM rainfall data
TRMM rainfall data maybe is the most accurate rainfall data derived from satellite measurements and a valuable source in regions with scarse weather-stations. It has a good spatial (0.25°) and temporal (daily) resolution and is available since 1998. However, downloading and processing may be a lot of work, if not scripted. The following script may be badly … Continue reading Automatically downloading and processing TRMM rainfall data